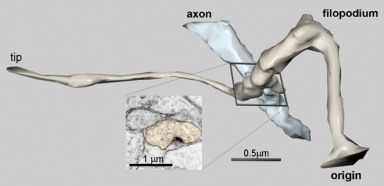
This image shows a 3D reconstruction of a small dendritic protrusion, a filopodium, and its synaptic interaction with an axon. The axon (blue) is less than 0.5 microns in diameter. (As a comparison, a human hair is at least 70 microns in diameter.) The filopodium (brown) is less than 0.1 microns in diameter in some places. The synaptic contact (inset) is only about 0.1 microns in its largest dimension. Since the wavelength of visible light is itself about 0.5 microns, the true dimensions of such small structures cannot be observed with light microscopy. More complex procedures are required to visualize them.
To visualize something this small, we begin with a small block of tissue containing the (even smaller) object. The tissue block is stained and embedded in epoxy. A small portion of the hardened block is shaved into a series of very thin sections, each about 50nm thick and 50 microns across, making a long ribbon of tissue sections.

Ribbons are transferred to specially prepared holders and stained with heavy metals. Then each section, or a portion of each section, is imaged at very high magnification using an electron microscope.
Rather than using light waves, the electron microscope passes electrons through the section. Photographs are made of the variations in electron density within the section. When the tissue has been properly stained, these patterns reveal the fine details of subcellular structures, down to a few nanometers.
The photographs of each section are digitized and then realigned into their original positions in the tissue volume using special-purpose computer software. After alignment of sections, the object of interest is then identified within this volume and a surface is generated by computer to represent its 3D spatial boundary.